Pluripotent stem cells (embryonic stem cells, ESC, and induced pluripotent stem cells, iPSC) can generate any of the hundreds of distinct cell types in the body. Thus, they are valuable tools for studying tissue formation and one day may provide a resource for cellular therapies.
We employ mouse and human ESC to study basic aspects of tissue development, and we derive personalized, patient-derived iPSC via reprogramming from somatic cells obtained from patients with variety of diseases. To reprogram somatic cells, we transfer genes responsible for the pluripotency of ESC into human fibroblasts or peripheral blood and derive cell lines that exhibit immortal self-renewal, multilineage differentiation potential in vitro, and teratoma formation in immunodeficient mice - currently the gold standard definition of human stem cell pluripotency. We have worked to improve the reprogramming method, to probe its mechanisms, and to assess how closely iPSC approximate embryo-derived stem cells.
We have generated iPSC from somatic cells obtained from patients with genetic immune deficiency, hemoglobin disorders like sickle cell anemia, and bone marrow failure syndromes including Fanconi anemia, dyskeratosis congenita, Shwachman-Diamond syndrome, Diamond-Blackfan anemia, and Pearson’s syndrome. These iPSC represent personalized human cell culture models that enable studies of fundamental disease mechanisms and in vitro chemical screens to identify drugs. We also use these disease models to deliver curative cell transplants after genome editing using for example the CRISPR/Cas system.
We have generated CellNet (http://cellnet.hms.harvard.edu/), a network biology-based computational platform that more accurately assesses the fidelity of cellular engineering than existing methodologies and generates hypotheses for improving cell derivations. Please see: "CellNet: Network Biology Applied to Stem Cell Engineering" Cahan et al., Cell 2014 August 14;158(4): 903-15; "Dissecting Engineered Cell Types and Enhancing Cell Fate Conversion via CellNet" Morris et al., Cell 2014 August 14:158(4):889-902.
Selected Publications:
Deconstructing transcriptional heterogeneity in pluripotent stem cells. Kumar RM, Cahan P, Shalek AK, Satija R, DaleyKeyser AJ, Li H, Zhang J, Pardee K, Gennert D, Trombetta JJ, Ferrante TC, Regev A, Daley GQ, Collins JJ. Nature. 2014 Dec 4;516(7529):56-61.
Dissecting Engineered Cell Types and Enhancing Cell Fate Conversion via CellNet. Samantha A. Morris, Patrick Cahan, Hu Li, Anna M. Zhao, Adrianna K. San Roman, Ramesh A. Shivdasani, James J. Collins, George Q. Daley. Cell 2014 August 14:158(4):889-902.
CellNet: Network Biology Applied to Stem Cell Engineering. Patrick Cahan, Hu Li, Samantha A. Morris, Edroaldo Lummertz da Rocha, George Q. Daley, James J. Collins. Cell 2014 August 14:158(4):903-15.
Development. A stem cell perspective on cellular engineering. Doulatov S, Daley GQ. Science. 2013 Nov 8;342(6159):700-2.
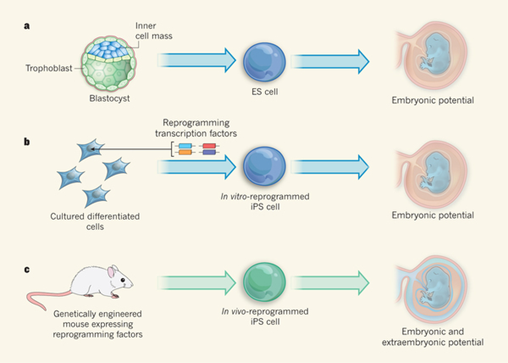
Stem cells: Reprogramming in situ. De Los Angeles A, Daley GQ. Nature. 2013 Oct 17;502(7471):309-10.
A chemical logic for reprogramming to pluripotency. De Los Angeles A, Daley GQ. Cell Res. 2013 Dec;23(12):1337-8.
Stem cell metabolism in tissue development and aging. Shyh-Chang N, Daley GQ, Cantley LC. Development. 2013 Jun;140(12):2535-47.
Origins and implications of pluripotent stem cell variability and heterogeneity. Cahan P, Daley GQ. Nat Rev Mol Cell Biol. 2013 Jun;14(6):357-68.
Reprogrammed cells for disease modeling and regenerative medicine. Cherry AB, Daley GQ. Annu Rev Med. 2013;64:277-90.
A blueprint for engineering cell fate: current technologies to reprogram cell identity. Morris SA, Daley GQ. Cell Res. 2013 Jan;23(1):33-48.
Cellular alchemy and the golden age of reprogramming. Daley GQ. Cell. 2012 Dec 7;151(6):1151-4.
